RFP - red florescent protein lab
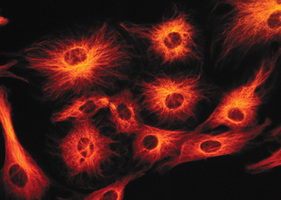
Purpose: make Red Florescent Protein (RFP) from jelly fish in bacteria
- Learn about steps done for genetic engineering
Materials/Procedure:
Lab 2A- Materials + procedure can be found in Amgen lab manual part 2A
Lab 4A- Materials + procedure can be found in Amgen lab manual part 4A
Lab 5A- Materials + procedure can be found in Amgen lab manual part 5A
Lab 6A- Materials + procedure can be found in Amgen lab manual part 6A
Experimental Overview:
Lab 2A: Verification of plasmid by restricting digest
- We cut a plasmid (circular DNA) with BaMH1 and HindIII so we are able to extract the segment of the RFP gene
Lab 4A- Verification of plasmid digest by electrophoresis
-To test that we cut the plasmid correctly, we used a process called electrophoresis, we ran the plasmid in a gel solution next to a DNA ladder. The DNA ladder is a ladder of bands that can correspond to other lanes of the gel to compare the size of the molecules. We completed this process correctly.
Lab 5A- Transformation of bacteria with recombinant plasmid
-We transformed the bacteria into a recombinant plasmid (man-made plasmid). We used restriction enzymes (molecular-scissors) to cut the plasmid and paste it to the gene of interest.
Lab 6A- Porification of RFP using chromotography
-we used a process called column chromotography to iscolate the RFP gene from everything else.
- this process includes running the gene through a series of hydrophilic beads, then extracting the gene with the buffer
Results:
- Learn about steps done for genetic engineering
Materials/Procedure:
Lab 2A- Materials + procedure can be found in Amgen lab manual part 2A
Lab 4A- Materials + procedure can be found in Amgen lab manual part 4A
Lab 5A- Materials + procedure can be found in Amgen lab manual part 5A
Lab 6A- Materials + procedure can be found in Amgen lab manual part 6A
Experimental Overview:
Lab 2A: Verification of plasmid by restricting digest
- We cut a plasmid (circular DNA) with BaMH1 and HindIII so we are able to extract the segment of the RFP gene
Lab 4A- Verification of plasmid digest by electrophoresis
-To test that we cut the plasmid correctly, we used a process called electrophoresis, we ran the plasmid in a gel solution next to a DNA ladder. The DNA ladder is a ladder of bands that can correspond to other lanes of the gel to compare the size of the molecules. We completed this process correctly.
Lab 5A- Transformation of bacteria with recombinant plasmid
-We transformed the bacteria into a recombinant plasmid (man-made plasmid). We used restriction enzymes (molecular-scissors) to cut the plasmid and paste it to the gene of interest.
Lab 6A- Porification of RFP using chromotography
-we used a process called column chromotography to iscolate the RFP gene from everything else.
- this process includes running the gene through a series of hydrophilic beads, then extracting the gene with the buffer
Results:

We extracted the RFP gene and ran it through a gel- the size of the DNA matched with the DNA ladder as we predicted.
2A: We verified that we had everything required for this lab and we are prepared for cutting the plasmid, at the end.
Questions:
1. Why does using two different enzymes to cut the plasmid prevent the plasmid from forming a circle without the inserted gene? Without the inserted gene after cutting a plasmid, would create an empty spot in the circle and cause the plasmid to never become a full circle.
2. What is the purpose of setting up a tube without BamH1 HindIII? - The purpose of setting up a tube without BamH1 HindIII is to create a control for the lab.
3. When do enzymes work best? - Enzymes work best at 37*C which is approximately because that is our body temperature and enzymes are in our body functioning normally. They are then frozen to stunt their growth.
4A: After running the gels it appeared that the cutting went good and the RFP gene and plasmid were the proper size off the DNA ladder.
Questions:
1. It was very useful to use loading dye in this lab to be able to see the plasmid and the RFP gene in the gel after running it.
2. We can predict the position of the R- and R+ tubes based on the DNA ladder because we do know their weights so we can gauge where they should fall on the ladder.
3. The DNA samples will be visible after the gel has run because the dyes will have dispersed onto the DNA and have made them visible.
5A: The bacteria appeared to have accepted the changes made to it and the correct bacteria lived based on the conditions that the bacteria was in.
Questions:
1. The P+ bacteria culture is treated differently because it has more plates than P- for the sake of comparing results. The P- culture is there to see the effects of LB and amp to the bacteria.
2. Cells need time to recover after the heat shock, otherwise they might die if they can't re-stabilize.
3. Cells are incubated at 37 degrees C because that is a normal temperature condition.
4. Aseptic is important in this lab because it keeps the bacteria from contaminating and becoming contaminated itself.
5. Most of my results except for the P+ only plate matched my predictions. I realized that the P+ was resistant to ara and amp, unlike I thought they were.
6A: The RFP easily binded to the beads to separate it and then we were able to obtain the RFP gene by itself with ease.
Questions:
1. You can determine where the rfp is in each step by watching for the red substance as it travels.
2. The supernatant is a bunch of clear cell nutrients and the pellet is compact bacterial cells with rfp.
3. The BB binds RFP to the resin bed, the WB washes away unwanted proteins, and EB knocks off RFP at the end.
4. A protein's conformation is important to it functioning because it needs to conform with its surroundings, otherwise it won't fit in and function with everything else like it should be.
2A: We verified that we had everything required for this lab and we are prepared for cutting the plasmid, at the end.
Questions:
1. Why does using two different enzymes to cut the plasmid prevent the plasmid from forming a circle without the inserted gene? Without the inserted gene after cutting a plasmid, would create an empty spot in the circle and cause the plasmid to never become a full circle.
2. What is the purpose of setting up a tube without BamH1 HindIII? - The purpose of setting up a tube without BamH1 HindIII is to create a control for the lab.
3. When do enzymes work best? - Enzymes work best at 37*C which is approximately because that is our body temperature and enzymes are in our body functioning normally. They are then frozen to stunt their growth.
4A: After running the gels it appeared that the cutting went good and the RFP gene and plasmid were the proper size off the DNA ladder.
Questions:
1. It was very useful to use loading dye in this lab to be able to see the plasmid and the RFP gene in the gel after running it.
2. We can predict the position of the R- and R+ tubes based on the DNA ladder because we do know their weights so we can gauge where they should fall on the ladder.
3. The DNA samples will be visible after the gel has run because the dyes will have dispersed onto the DNA and have made them visible.
5A: The bacteria appeared to have accepted the changes made to it and the correct bacteria lived based on the conditions that the bacteria was in.
Questions:
1. The P+ bacteria culture is treated differently because it has more plates than P- for the sake of comparing results. The P- culture is there to see the effects of LB and amp to the bacteria.
2. Cells need time to recover after the heat shock, otherwise they might die if they can't re-stabilize.
3. Cells are incubated at 37 degrees C because that is a normal temperature condition.
4. Aseptic is important in this lab because it keeps the bacteria from contaminating and becoming contaminated itself.
5. Most of my results except for the P+ only plate matched my predictions. I realized that the P+ was resistant to ara and amp, unlike I thought they were.
6A: The RFP easily binded to the beads to separate it and then we were able to obtain the RFP gene by itself with ease.
Questions:
1. You can determine where the rfp is in each step by watching for the red substance as it travels.
2. The supernatant is a bunch of clear cell nutrients and the pellet is compact bacterial cells with rfp.
3. The BB binds RFP to the resin bed, the WB washes away unwanted proteins, and EB knocks off RFP at the end.
4. A protein's conformation is important to it functioning because it needs to conform with its surroundings, otherwise it won't fit in and function with everything else like it should be.

This is the picture of the RFP gel after we performed the column chromatography.
Data/Analysis:
Our red conies didn't show up so our experiment went wrong at some point. Maybe we cross-contaminated one of the buffers but I don't think so. Our R+ and R- met our predictions so we didn't go wrong there, but I would further like to research what we did wrong.
Reflection: Overall this lab was really fun handling atomic level molecules. I learned alot by hands on experience with the hardware in the lab such as the mini centrifuge and with running the gel. This lab experience will help us in the field of biotech for more complex labs in the future. Thank you Stem Marin for supporting our program!
Our red conies didn't show up so our experiment went wrong at some point. Maybe we cross-contaminated one of the buffers but I don't think so. Our R+ and R- met our predictions so we didn't go wrong there, but I would further like to research what we did wrong.
Reflection: Overall this lab was really fun handling atomic level molecules. I learned alot by hands on experience with the hardware in the lab such as the mini centrifuge and with running the gel. This lab experience will help us in the field of biotech for more complex labs in the future. Thank you Stem Marin for supporting our program!
